Social network analysis with NetworkX
Introduction
In this tutorial, we will show you how to perform a simple network analysis with the NetworkX library and data stored in Memgraph. You will also acquire a basic understanding of query modules, an easy method for extending the query language with user-written procedures.
To get started, sign up to Memgraph Cloud, create a new instance and connect to it using in-browser Memgraph Lab. If you require help, check out the documentation on Memgraph Cloud.
You can also install Memgraph using the memgraph-platform image by following
the installment instructions for your OS. Once
Memgraph is up and running, connect to it using Memgraph Lab, a visual user
interface that you can also use from your browser at
http://localhost:3000 or download as an
application.
Data model
We are going to use the Karate Club graph, a network of friendships between 34
members of a karate club at a US university, as described by Wayne Zachary in
1977. It is a very popular data set in social network analysis and is very often
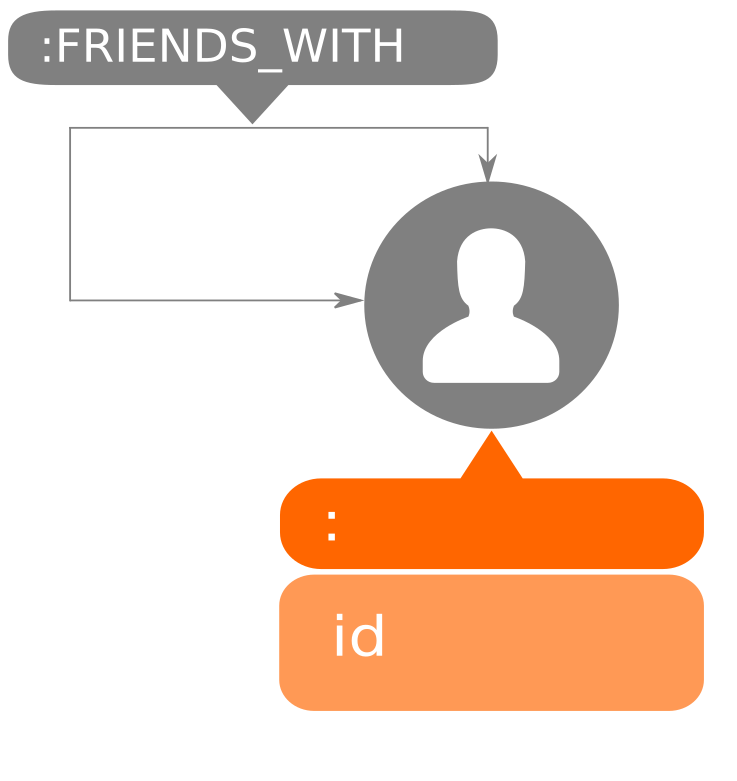
referenced in such tutorials. The nodes in the graph represent the members while
the relationships between them are of type FRIENDS_WITH. You can differentiate
the nodes by using their unique id property.
Import the dataset
To import the dataset, open Memgraph Lab and navigate to the Datasets tab in
the sidebar. From there, load the Karate club friendship network dataset, wait
for the import to finish, move to the Query tab and continue with the
tutorial.
Use existing NetworkX algorithms
Execute the following command to get all the relationships inside our network:
MATCH (s)-[r]-(t)
RETURN s, r, t;
Now we have a better overview of what we are dealing with, so it’s time to get some useful information about the network.
To analyze the network we will use the built-in procedure analyze() from the
graph_analyzer query module. This module utilizes the NetworkX library to
retrieve graph information. Run the following query:
CALL graph_analyzer.analyze() YIELD *;
You will get details about the graph, such as the number of nodes, edges, bridges and many more.
Betweenness centrality
Now let's try to find the betweenness centrality of a node, i.e. the number of times a node acts as a bridge along the shortest path between two other nodes. Run the following query:
CALL nxalg.betweenness_centrality() YIELD *;
The procedure betweenness_centrality() is one of over 70 algorithms available
in the nxalg module.
The result should be:
+--------------+--------------+
| betweenness | node |
+--------------+--------------+
| 0 | ({id: "0"}) |
| 0.000473485 | ({id: "1"}) |
| 0.0083649 | ({id: "2"}) |
| 0.00189394 | ({id: "3"}) |
| 0 | ({id: "4"}) |
| 0.000473485 | ({id: "5"}) |
| ... | ... |
Link prediction
A very common problem in network analysis is link prediction. The algorithm
predicts which new interactions among the network members are likely to occur in
the near future. One way of predicting these links is by measuring the
“proximity” of nodes in a network. This can be done by using the Jaccard
coefficient. Let's try running the algorithm on a node with the id 13 and
ordering the results descending by the value of the coefficient:
CALL nxalg.jaccard_coefficient()
YIELD *
WITH u, v, coef
WHERE u.id = '13'
RETURN u, v, coef
ORDER BY coef DESC;
The results are:
+--------------+--------------+--------------+
| u | v | coef |
+--------------+--------------+--------------+
| ({id: "13"}) | ({id: "19"}) | 0.6 |
| ({id: "13"}) | ({id: "17"}) | 0.4 |
| ({id: "13"}) | ({id: "21"}) | 0.4 |
| ({id: "13"}) | ({id: "28"}) | 0.333333 |
| ({id: "13"}) | ({id: "30"}) | 0.285714 |
| ({id: "13"}) | ({id: "27"}) | 0.285714 |
| ({id: "13"}) | ({id: "31"}) | 0.222222 |
| ({id: "13"}) | ({id: "15"}) | 0.166667 |
| ({id: "13"}) | ({id: "14"}) | 0.166667 |
| ({id: "13"}) | ({id: "18"}) | 0.166667 |
| ({id: "13"}) | ({id: "20"}) | 0.166667 |
| ({id: "13"}) | ({id: "22"}) | 0.166667 |
| ({id: "13"}) | ({id: "26"}) | 0.166667 |
| ({id: "13"}) | ({id: "32"}) | 0.133333 |
| ({id: "13"}) | ({id: "29"}) | 0.125 |
| ({id: "13"}) | ({id: "23"}) | 0.111111 |
| ({id: "13"}) | ({id: "25"}) | 0 |
| ({id: "13"}) | ({id: "24"}) | 0 |
| ({id: "13"}) | ({id: "16"}) | 0 |
+--------------+--------------+--------------+
Add new NetworkX algorithms as query modules
Memgraph comes with over 70 NetworkX algorithms, but if the algorithm you require is missing, you can add it yourself as a query module.
Let's create a custom query module!
Community detection algorithm
Detecting communities in a network is a very common problem. Therefore, we need community detection algorithms that can partition the network into multiple communities. Let's create our own module that accomplishes this task.
Go to the Query Modules section in Memgraph Lab and click on the + New Module button. Give it a name, such as communities and Create it. A new query module will be created with example procedures. Feel free to erase them and copy the following code into it:
import mgp
import networkx as nx
from networkx.algorithms import community
from mgp_networkx import MemgraphDiGraph
@mgp.read_proc
def detect(
ctx: mgp.ProcCtx
) -> mgp.Record(communities=mgp.List[mgp.List[mgp.Vertex]]):
networkxGraph = nx.DiGraph(MemgraphDiGraph(ctx=ctx))
communities_generator = community.girvan_newman(networkxGraph)
return mgp.Record(communities=[
list(s) for s in next(communities_generator)])
Click Save and you should be able to see the procedure and its signature as
Detected procedures & transformations. This query module with the procedure
detect() utilizes the Girvan–Newman method to find communities in a graph.
Save and close the window then move to the Query Execution section to use the procedure.
Call the query module
Let's call the custom query module with Cypher:
CALL communities.detect()
YIELD communities
UNWIND communities AS community
RETURN community;
The resulting communities are:
+-------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------+
| community |
+-------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------+
| [({id: "0"}), ({id: "1"}), ({id: "3"}), ({id: "4"}), ({id: "5"}), ({id: "6"}), ({id: "7"}), ({id: "10"}), ({id: "11"}), ({id: "12"}), ({id: "13"}), ({id: "16"}), ({id: "17"}), ({id: "19"}), ({id: "21"})] |
| [({id: "2"}), ({id: "8"}), ({id: "9"}), ({id: "14"}), ({id: "15"}), ({id: "18"}), ({id: "20"}), ({id: "22"}), ({id: "23"}), ({id: "24"}), ({id: "25"}), ({id: "26"}), ({id: "27"}), ({id: "28"}), ({id: "29"}), ({id: "30"}), ({id: "31"}), ({id: "32"}), ({id: "33"})] |
+-------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------+
Further reading
If you want to find out more about query modules, take a look at our guide on how to create your own: Implement custom query modules.
You can also visit our NetworkX Reference guide to find out which NetworkX algorithms are already available in Memgraph.